loading…
ClaudeR
БесплатноConnect RStudio directly to Claude Desktop, Claude Code, Codex, Gemini CLI, or any other MCP based AI assistant for interactive coding and data analysis.
Описание
Connect RStudio directly to Claude Desktop, Claude Code, Codex, Gemini CLI, or any other MCP based AI assistant for interactive coding and data analysis.
README

ClaudeR - The Modern Researcher's Toolkit
Connect RStudio to Claude Code, Codex, Gemini CLI, or any MCP-based LLM agent for interactive coding, multi-agent orchestration, automated manuscript auditing, and data annotation.
ClaudeR is an R package that forges a direct link between RStudio and MCP configured LLM agents like Claude Code or Codex. This allows interactive coding sessions where the agent can execute code in your active RStudio environment so it can see the executed code and any generated plots in real-time. If you need help editing a script, a quick analysis done, or an LLM to audit your statistical claims against any manuscript before submission: ClaudeR has got your back.
This package, additionally, allows multiple agents to work on one script, or it can make multiple RStudio windows siloed so multiple agents can operate independently on different datasets. It's also compatible with Cursor and any service that support MCP servers.
Quick Start
# Install
if (!require("devtools")) install.packages("devtools")
devtools::install_github("IMNMV/ClaudeR")
# Set up your AI tool
library(ClaudeR)
install_clauder() # For Claude Desktop / Cursor
install_cli(tools = "claude") # For Claude Code CLI
# Start the server in RStudio
claudeAddin()
AI agents: See llms-install.md for automated setup instructions.
Recent Updates (click to expand)
- AI-Driven Data Annotation. Five MCP tools (
load_annotation_data,annotate,run_annotation_job,get_annotation_job_status,cancel_annotation_job) let an agent label a CSV dataset row by row without writing any code. Two modes: interactive (context accumulates across rows) or subprocess-per-row viarun_annotation_jobfor full row isolation usingclaudeorcodex. Codex jobs accept areasoning_effortparameter. The original file is never modified and sessions resume automatically if interrupted. - Multi-Agent Coordination Protocol. Built-in protocol for multiple agents sharing one RStudio session. Agents negotiate through a shared message board in the R environment, agree on a task plan, claim tasks before working, and cross-check each other's output. Load it with
multi_agent_prompt(). verify_referencestool. Extracts DOIs from a manuscript's bibliography, queries the CrossRef API for each, and returns metadata (title, authors, year, journal) for comparison against manuscript claims. Non-resolving DOIs, metadata mismatches, and references without DOIs are flagged. Works standalone ("check my references") or as Pass 4 of Reviewer Zero.- R Best Practices Protocol. Built-in statistical analysis protocol covering EDA, assumption checking, model building, diagnostics, multiple-corrections, and reporting. Load it with
r_best_practices_prompt()or tell the agent to read it. - Reviewer Zero: Automated Academic Audits. Now a 4-pass protocol for AI-driven manuscript verification. The agent extracts every statistical and methodological claim, verifies its extraction, recomputes values against the author's R code, and checks references via CrossRef. Methodological claims (e.g., "zero variance made testing impossible") are tested directly rather than accepted at face value. Run
reviewer_zero_prompt()to get the full protocol. clean_error_logtool. Point the agent at a session log and it will parse every code block, find errors, check whether a fix follows each one, then strip the error blocks and any duplicate code that preceded them. The result is a clean log with only the working code. Accepts an optionaloutput_pathto write to a separate file instead of overwriting the original.- Persistent server across UI restarts. Closing the Shiny addin (console stop or Done button) no longer kills the MCP server. Re-running
claudeAddin()reconnects to the still-running server with the correct port, session name, and execution count. Only clicking "Stop Server" in the UI actually stops the server. - Descriptive log filenames. Log files now include the session name, port, and timestamp:
clauder_default_8787_20260301_143022.R. A new log file is created each time you click Start Server. All subsequent code execution appends to that file. - Viewer content capture &
insert_texttool. Two new tools:get_viewer_contentreads HTML from interactive widgets (plotly, DT, leaflet) with pagination so agents can inspect htmlwidget output without blowing up context.insert_textinserts text at the cursor position or a specific line/column in the active document. During agent execution, htmlwidgets open in the browser instead of stealing the Shiny addin's viewer pane. - Multi-session routing fix. Agents now prefer the session named "default" when multiple sessions are active, preventing misrouting caused by non-deterministic discovery order. Once bound, agents stay sticky to their session. Non-default agents should call
connect_sessionto target a specific session. - Reproducibility metadata in logs. When logging is enabled, each new session log starts with a header containing the date, working directory, and full
sessionInfo()output (R version, platform, attached packages). Anyone who receives the log can see exactly what environment the code ran in. - Export clean script. Click "Export Clean Script" in the Shiny addin to strip all timestamps, agent labels, and log headers from a session log, producing a runnable
.Rfile with just the code. Error blocks are preserved as comments. Also available programmatically viaexport_log_as_script(). - PyPI package (
clauder-mcp). The Python MCP bridge is now available as a standalone package on PyPI. Run it withuvx clauder-mcpfor zero-config setup with no Python path or pip install needed. The installers (install_cli()andinstall_clauder()) default to uvx, with ause_uvx = FALSEfallback for legacy setups. read_filetool. Agents can now read any text file from disk (.R, .qmd, .csv, .log, etc.) without it being open in RStudio. Enables session continuity workflows: point an agent at a previous log file and tell it to pick up where the last session left off.- Codex CLI support.
install_cli(tools = "codex")generates the setup command for OpenAI Codex. Codex joins Claude Code and Gemini as a supported CLI agent. - Multi-agent orchestration. Run multiple AI agents on the same R session or spread them across separate RStudio windows. Each agent gets a unique ID on startup. Console output, log files, and execution history are all attributed per agent, so you always know who did what. On its very first tool call, each agent receives a context briefing with its own ID, any other agents active on the session, and the log file path, giving it full awareness of the shared environment without any manual setup. Agents can call
get_session_historyto review what other agents have done, or read the shared log file directly. The Shiny viewer tracks connected agents in real-time. - Session discovery. Each RStudio session writes a discovery file to
~/.claude_r_sessions/on startup. AI agents find sessions automatically with no hardcoded ports. Name your sessions (e.g. "analysis", "modeling") and run them on different ports. When multiple sessions exist, agents automatically route to the session named "default". Non-default agents should callconnect_sessionto bind to their target session. Single-session setups work with zero config. - Redesigned Shiny viewer. Cleaner UI with grouped panels for Session, Agents, Logging, and Advanced settings. Shows connected agents and execution count in real-time. Click the
?button for a built-in guide on multi-session setup and agent identity. - Non-blocking async execution.
execute_r_asyncnow runs long-running code in a separate R process viacallr, keeping the main session fully responsive. Other agents can continue working while a job runs. The agent writes self-contained code (explicitly saving/loading data viasaveRDS), submits it, and polls withget_async_result. No environment copying and no memory doubling, only the data the job needs gets serialized. - Stale plot detection. Fixed a bug where the last generated plot image would persist and re-appear on every subsequent
execute_rcall, even when no new plot was created. - Reduced plot token usage. Plot capture now uses smaller dimensions (600x400, dpi 100) to reduce base64 image size and avoid token overflow errors.
- MCP tool annotations. All tools now include
readOnlyHint,destructiveHint, andidempotentHintannotations per the current MCP spec. - Hardened string escaping.
escape_r_stringnow handles backticks, carriage returns, tabs, and null bytes. Applied to task tool inputs to prevent injection. - Fixed
install_cli()command syntax. Updated to use--transport stdioflag and--separator for current Claude Code CLI. Now removes stale MCP registrations before adding fresh ones, preventing issues when upgrading R versions.
Demo
| Single agent via Claude Desktop App | Multi-agent: Codex + Claude Code via CLI | GPT 5.4 Codex: Data analysis + Quarto report |
|---|---|---|
| Single Agent Demo | Multi-Agent Demo | Codex Quarto Demo |
Table of Contents
- Quick Start
- Features
- How It Works
- Reviewer Zero
- R Best Practices Protocol
- Multi-Agent Coordination Protocol
- CLI Integration
- Security Restrictions
- Installation
- Usage
- Logging Options
- Example Interactions
- Important Notes
- Troubleshooting
- Limitations
- License
- Contributing
Features
ClaudeR empowers your AI assistant with a suite of tools to interact with your R environment:
execute_r: Execute R code and return the output.execute_r_with_plot: Execute R code that generates a plot that the model can see.execute_r_async: Execute long-running R code asynchronously (>25 seconds). Returns a job ID for polling.get_async_result: Poll for the result of an async job. Includes a built-in delay to throttle polling.list_sessions: List all active RStudio sessions the agent can connect to.connect_session: Connect to a specific RStudio session by name for multi-session workflows.get_session_history: View execution history filtered by agent ID.read_file: Read any text file from disk (.R, .qmd, .csv, .log, etc.) without needing it open in RStudio. Supportsstart_line/end_linepagination for large files.get_active_document: Get the content of the active document in RStudio.get_r_info: Get information about the R environment.modify_code_section: Modify a specific section of code in the active document.insert_text: Insert text at the current cursor position or a specific line/column in the active document.get_viewer_content: Read HTML content from the viewer pane (plotly, DT, leaflet widgets) with pagination support.clean_error_log: Clean a session log by removing error blocks and their duplicate predecessors, leaving only working code and the fixes that followed.search_project_code: Search for a regex pattern across project source files (.R, .Rmd, .qmd). Returns file, line number, and snippet.probe_scripts: Source R scripts in a clean background session and report what objects are created (names, classes, dimensions) without affecting your main session.verify_references: Verify academic references by extracting DOIs and checking them against the CrossRef API. Returns metadata (title, authors, year, journal) for comparison. References without DOIs are flagged for manual web search.create_task_list: Generate a task list based on your prompt to prevent omissions in long-context tasks.update_task_status: Track progress for each task in the generated list.load_annotation_data: Load a CSV for annotation. Creates a working copy, parses the_schemacolumn, and displays the first unannotated row. Resumable if interrupted.annotate: Annotate the current row, validate against the schema, save immediately, and auto-load the next row.run_annotation_job: Annotate a full CSV in the background using a fresh subprocess per row with no context bleed between rows. Supportsclaudeandcodex. Accepts areasoning_effortparameter (low,medium,high) when using Codex.get_annotation_job_status: Check progress of a running or completed annotation job.cancel_annotation_job: Cancel a running background annotation job. Rows completed before cancellation are preserved in the output file.
With these tools, you can:
- Direct Code Execution: The AI can write, execute, and install packages in your active RStudio session.
- Feedback & Assistance: Get explanations of your R scripts or request edits at specific lines.
- Visualization: The AI can generate, view, and refine plots and visualizations.
- Data Analysis: Let the AI analyze your datasets and iteratively provide insights.
- Multi-Agent Workflows: Run Claude Desktop, Claude Code, and Gemini CLI on the same R session simultaneously. Each agent is uniquely identified, and they can see each other's work through shared history and log files.
- Long-Running Analysis: Async execution handles model fitting, simulations, and large data processing without timing out.
- Code Logging: Save all code executed by the AI to log files for future reference. Every entry is tagged with the agent that ran it.
- Console Printing: Print the AI's code to the console before execution.
- Environment Integration: The AI can access variables and functions in your R environment.
- Dynamic Summaries: Summaries can dynamically pull results from objects and data frames to safeguard against hallucinations.
- Quarto Renders: The AI can create and render Quarto presentations. For best results, ask for a .qmd file and for it to be rendered in HTML when it's finished.
- Reviewer Zero: A built-in protocol for automated academic auditing. The AI reads a manuscript block-by-block, extracts every statistical claim into a registry, verifies its extraction, then recomputes each claim against the author's R code. Run
reviewer_zero_prompt()for the full protocol. See the Reviewer Zero section below. - Data Annotation: Label CSV datasets row by row using the built-in annotation tools. Define the annotation schema in a
_schemacolumn, rundata_annotation_prompt()to get the protocol, and annotate interactively or in fully isolated subprocess-per-row mode withrun_annotation_job.
Reviewer Zero: Automated Academic Audits
ClaudeR includes a built-in protocol for AI-driven technical review of academic manuscripts. The AI acts as "Reviewer Zero": systematically verifying that every p-value, coefficient, and confidence interval in your paper matches the code that produced it.
How it works (4-pass protocol):
- Extract: The AI reads your manuscript block-by-block using paginated
read_file, extracting every quantitative and methodological claim into a structured registry (a data.frame visible in your RStudio Environment pane). - Verify: The AI re-reads the source lines for each claim to confirm it didn't misread values. No code runs until every claim is verified.
- Recompute: The AI uses
search_project_codeandprobe_scriptsto locate the relevant R scripts, thenexecute_rto rerun the analyses and compare recomputed values against the manuscript. Methodological claims (e.g., "zero variance made testing impossible") are tested directly rather than accepted at face value. - References: The AI uses
verify_referencesto extract DOIs from the bibliography and check each against the CrossRef API. Metadata mismatches, non-resolving DOIs, and references without DOIs are flagged. In-text citations are cross-checked against the bibliography.
To get started:
# Print the full protocol prompt to give to your AI agent
reviewer_zero_prompt()
The protocol works with .docx, .pdf, .qmd, .Rmd, .tex, or plain text manuscripts and supports multi-script R projects.
R Best Practices Protocol
ClaudeR comes with a built-in statistical analysis protocol inspired by the modeling workflows I learned from my statistics courses and refined through oof moments from using AI agents in real statistical work. The goal is to steer models with natural language to reproducible, theory-driven analysis which covers EDA, assumption checking, model building, diagnostics, multiple-corrections, and reporting.
# Print the full protocol to give to your AI agent
r_best_practices_prompt()
You can also just tell the agent to run ClaudeR::r_best_practices_prompt() and it will read the protocol itself.
Multi-Agent Coordination Protocol
When two or more agents share the same RStudio session, they need a way to divide work without stepping on each other. The multi-agent protocol handles this with a structured workflow: agents check in by reading the session log, the first agent creates a task plan, agents claim tasks before starting them, and they cross-check each other's work when done.
# Print the full protocol to give to your AI agents
multi_agent_prompt()
You can also just tell the agents to run ClaudeR::multi_agent_prompt() and they will read the protocol themselves.
AI-Driven Data Annotation
ClaudeR includes a purpose-built annotation workflow for labelling CSV datasets with an AI agent. The agent works through the dataset row by row using two dedicated MCP tools with no code required on the agent's end.
CSV format: add a _schema column to your file and define the annotation fields in the first row using a simple type syntax:
text,label,confidence,_schema
"Some text","","","label:choice[positive,negative,neutral];confidence:float[0,1]"
"More text","","",""
Supported types: choice[a,b,c], float[min,max], int[min,max], bool, text
Running an annotation session:
# Print the full protocol to give to your agent
data_annotation_prompt()
Or tell the agent to run ClaudeR::data_annotation_prompt() and it will read the protocol itself. The agent then calls load_annotation_data to start and annotate to label each row. The original file is never modified and sessions are automatically resumable if interrupted.
Two annotation modes are available: The default load_annotation_data + annotate flow runs inside the agent's existing conversation where context accumulates across rows, which can be useful for consistency but may introduce anchoring on long datasets. For full row isolation, use run_annotation_job instead: it spawns a fresh claude or codex subprocess per row so each annotation is made with zero memory of prior rows.
Mode 1: Full context (interactive):
You are annotating a dataset. Your only job is to call annotation tools. Do not write code or use any other tools.
Step 1: Call load_annotation_data with:
- csv_path: /path/to/your/file.csv
Step 2: For each row displayed, call annotate with the fields defined in the schema.
Step 3: After each annotate call, the next row loads automatically. Keep annotating until you see "Annotation complete."
If you get a validation error, read it carefully and call annotate again with corrected values.
Mode 2: Isolated context (subprocess per row):
You are annotating a dataset.
Step 1: Call run_annotation_job with:
- csv_path: /path/to/your/file.csv
- tool: claude # or "codex"
- reasoning_effort: high # codex only: low | medium | high
Step 2: Once you have the job ID, periodically call get_annotation_job_status with that ID to check progress.
To stop early, call cancel_annotation_job with the job ID. Rows already annotated are preserved.
That's it. The annotation runs automatically in the background. Do not call any other tools unless checking status.
How It Works
ClaudeR uses the Model Context Protocol (MCP) to create a bidirectional connection between an AI assistant and your RStudio environment. MCP is an open protocol from Anthropic that allows the AI to safely interact with local tools and data.
Here's the workflow:
- The Python MCP server acts as a bridge.
- The AI sends a code execution request to the MCP server.
- The server forwards the request to the R add-in running in RStudio.
- The code executes in your R session, and the results are sent back to the AI.
This architecture ensures that the AI can only perform approved operations through well-defined interfaces, keeping you in complete control of your R environment.
CLI Integration
ClaudeR now supports command-line interface (CLI) tools like the Claude Code CLI, the OpenAI Codex CLI, and the Google Gemini CLI. This is ideal for developers who prefer a terminal-based workflow, allowing you to interact with your AI assistant directly from the command line while maintaining a live connection to your RStudio session.
Security Model
ClaudeR is a supervised power tool. The agent executes R code in your live RStudio session, the same session where your data and variables live. You should review what it does, just as you would review a colleague's code before running it.
What ClaudeR blocks
- System commands:
system(),system2(),shell(), and related calls are blocked to prevent the agent from reaching outside R. - File deletion:
unlink(),file.remove(), and shell commands containingrmare prohibited. - Error feedback: Blocked operations return a clear error message explaining why.
What ClaudeR does NOT restrict
The agent can still read files, install packages, create/overwrite objects in your environment, and consume compute resources. These are necessary for the agent to be useful, but they mean you should:
- Use logging (enabled by default) so you have a full record of every line the agent executed and which agent ran it.
- Work in a project directory to limit what the agent can see.
- Review before trusting: especially for Reviewer Zero audits, treat the output as a draft review that should be verified.
These restrictions only apply to code executed by the AI. Your manually executed R code is not affected.
Installation
Step 1: Install ClaudeR from GitHub
Run this command in your RStudio console:
if (!require("devtools")) install.packages("devtools")
devtools::install_github("IMNMV/ClaudeR")
Step 2: Run the Correct Installer
Choose the option that matches your workflow.
Option A: For Desktop Apps (Claude Desktop / Cursor)
This function configures the MCP config file automatically for desktop applications. By default it uses uvx to run the clauder-mcp PyPI package, which handles all Python dependencies automatically.
# Load the package
library(ClaudeR)
# Run the installer for Claude Desktop
install_clauder()
# Optional: For Cursor users
# install_clauder(for_cursor = TRUE)
For users who cannot use uvx (e.g. restricted environments), fall back to the legacy Python path method:
library(ClaudeR)
install_clauder(use_uvx = FALSE, python_path = "/path/to/your/python")
Option B: For CLI Tools (Claude Code / Codex / Gemini)
This non-interactive function generates the exact command or JSON configuration needed for your CLI tool.
library(ClaudeR)
# For Claude Code CLI
install_cli(tools = "claude")
# For OpenAI Codex CLI
install_cli(tools = "codex")
# For Google Gemini CLI
install_cli(tools = "gemini")
For users who cannot use uvx, fall back to the legacy Python path method:
install_cli(tools = "claude", use_uvx = FALSE, python_path = "/path/to/my/python")
After running the function, you must manually apply the configuration:
- For Claude / Codex: Copy the command printed in the R console and run it in your terminal.
- For Gemini: Copy the generated JSON and manually add it to your
gemini.jsonsettings file.
After setup, quit and restart any active Desktop Apps or terminal sessions for the new settings to load.
Note: If you upgrade R versions, re-run
install_cli()orinstall_clauder()to update the MCP server path. The CLI installer automatically removes stale registrations before adding fresh ones.
Usage
Part 1: In RStudio
For all workflows, you must first start the ClaudeR server from RStudio.
library(ClaudeR)
claudeAddin()
The ClaudeR add-in will appear in your RStudio Viewer pane. Click "Start Server". Keep this window active while using your preferred tool.
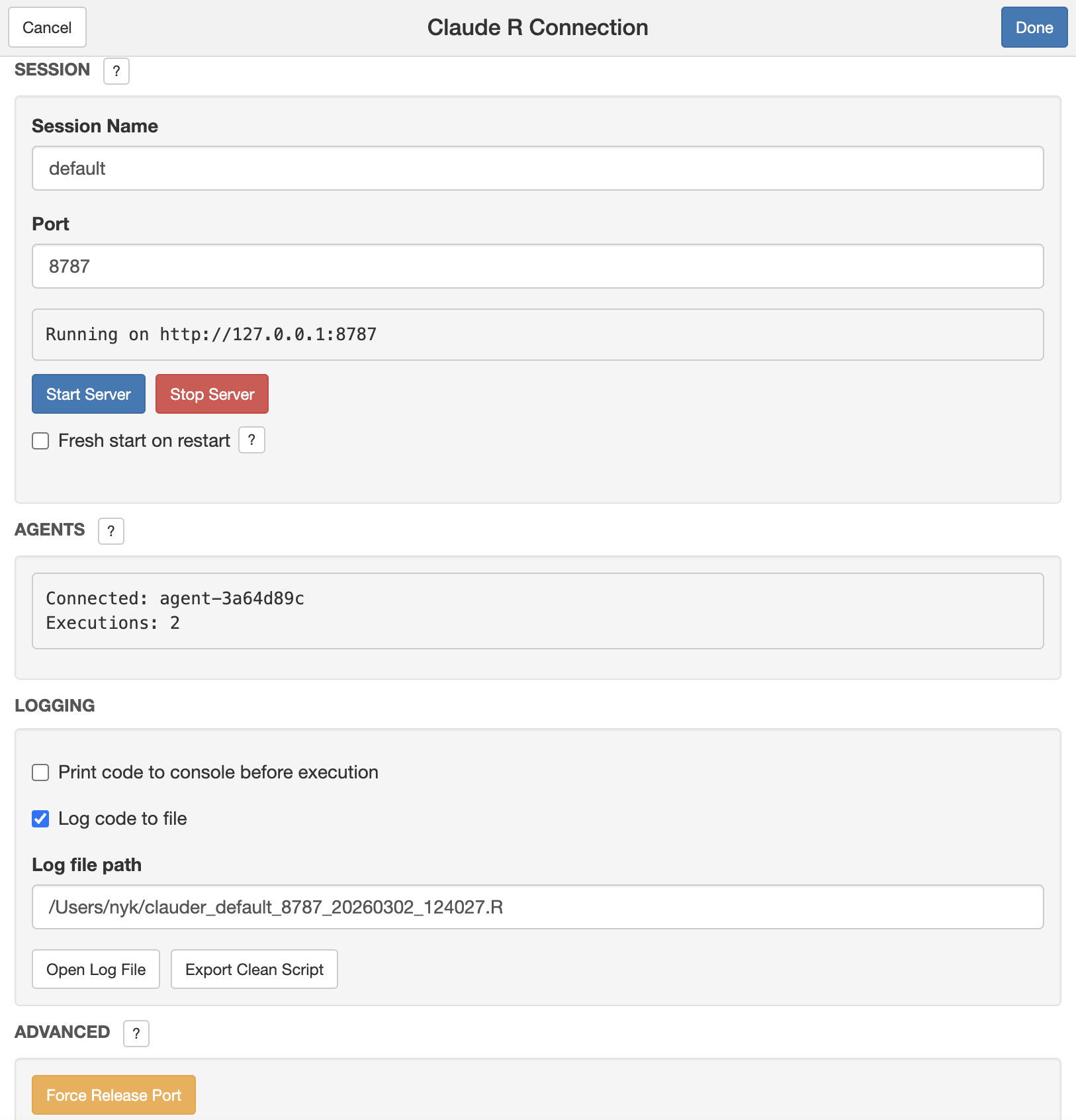
Part 2: In Your AI Tool
- For Desktop Apps: Open the Claude Desktop App or Cursor and begin your session.
- For CLI Tools: Open your terminal and use the
claudeorgeminicommands to start interacting with your AI assistant.
Note: You can regain console/active document control by clicking the stop button in the RStudio console. This closes the Shiny UI but the MCP server keeps running in the background and your AI agents stay connected. Re-run
claudeAddin()to bring the viewer pane back with the same server state (port, session name, execution count). To fully stop the server, click "Stop Server" in the UI before closing.
Logging Options
- Print Code to Console: See the AI's code in your R console before it runs. The code will be preceded by the header:
### LLM [agent-id] executing the following code ###. - Log Code to File: Save all executed code to a log file. Each entry is tagged with the agent ID that executed it, so you can trace which AI agent ran what.
- Custom Log Path: Specify a custom location for log files.
- Descriptive Filenames: Log files are named
clauder_<session>_<port>_<timestamp>.R(e.g.,clauder_default_8787_20260301_143022.R) so you can tell at a glance which session produced which log. - Reproducibility Header: Each log starts with a header containing the date, working directory, and full
sessionInfo()output (R version, platform, attached/loaded packages). This makes logs self-documenting for reproducibility. - Export Clean Script: Click "Export Clean Script" in the logging panel to produce a runnable
.Rfile stripped of all timestamps and log headers. Error blocks are kept as comments so you can see what went wrong. Also callable from the console withexport_log_as_script().
A new log file is created each time you click Start Server. All code executed by agents appends to that file until you stop and start the server again.
Example Interactions
- "I have a dataset named
datain my environment. Perform exploratory data analysis on it." - "Load the
mtcarsdataset and create a scatterplot ofmpgvs.hpwith a trend line." - "Fit a linear model to predict
mpgbased onwtandhp." - "Generate a correlation matrix for the
irisdataset and visualize it." - "I have a qmd file active. Please make a nice quarto presentation on gradient descent. The audience is very technical. Make sure it looks smooth. Save the presentation in /Users/nyk/QuartoDocs/"
If you can do it with R, your AI assistant can too.
Important Notes
- Session Persistence: Variables, data, and functions created by the AI remain in your R session.
- Code Visibility: By default, the AI's code is printed to your console.
- Port Configuration: The default port is
8787, but you can change it if needed. - Package Installation: The AI can install packages. Use clear prompts to guide its behavior.
Troubleshooting
- Connection Issues:
- Ensure your AI tool is configured correctly.
- Verify the Python path in your
configor CLI command. - Make sure the server is running in the add-in.
- Restart RStudio if the port is in use.
- Python Dependency Issues:
could not find function install_clauder: Restart your R session (Session -> Restart R) and try again.- MCP Server Failed to Start: If using
uvx, ensureuvis installed (curl -LsSf https://astral.sh/uv/install.sh | sh). If using the legacy method, this usually means the wrong Python environment was detected. Re-run the installer with the correctpython_pathor switch touse_uvx = TRUE.
- AI Can't See Results:
- Ensure the add-in window is open and the server is running.
- Plots Not Displaying:
- Instruct the AI to wrap plot objects in
print()(e.g.,print(my_plot)). - Tell the AI to use the
execute_r_with_plotfunction.
- Instruct the AI to wrap plot objects in
- Long-Running Code Timing Out:
- Ask the AI to use
execute_r_asyncfor code that takes longer than 25 seconds. - The AI will automatically poll for results using
get_async_result. - Async jobs run in a separate R process via
callrand do not have access to your main session's environment. The AI must write self-contained code that usessaveRDS()to pass data in and write results out, then loads them back into the main session after the job completes.
- Ask the AI to use
- Server Restart Issues:
- If you see an "address already in use" error after restarting the server, it's a UI bug. The server is still active. If you encounter connection issues, switch the port number in the Viewer Pane or restart RStudio.
- If the AI still can't connect, click "Force Release Port" under the Advanced section. This force-kills whatever process is holding the port so you can start fresh.
- Stale MCP Path After R Upgrade:
- If tools stop working after upgrading R, re-run
install_cli()orinstall_clauder()to update the script path.
- If tools stop working after upgrading R, re-run
Limitations
- Each R session can connect to one Claude Desktop/Cursor app at a time. However, multiple CLI agents (Claude Code, Gemini CLI) can share the same session alongside a Desktop app. To isolate agents, run separate RStudio windows with different session names and ports.
- You can stop the connection to the Shiny UI by clicking the Stop button in the console to make changes alongside the AI, but to stop the connection you will need to restart the RSession.
- R is single-threaded, but async jobs run in a separate process via
callrso the main session stays responsive. The background process does not share the main session's environment, so async code must be self-contained.
License
This project is licensed under the MIT License. See the LICENSE file for details.
Contributing
Contributions are welcome! Please feel free to submit a Pull Request.
Как установить
Добавь это в claude_desktop_config.json и перезапусти Claude Desktop.
{
"mcpServers": {
"clauder": {
"command": "npx",
"args": []
}
}
}Похожие MCP
GitHub
PRs, issues, code search, CI status
Supabase
Database, auth and storage
Everything
Reference / test server with prompts, resources, and tools.
Filesystem
Secure file operations with configurable access controls.





